Background
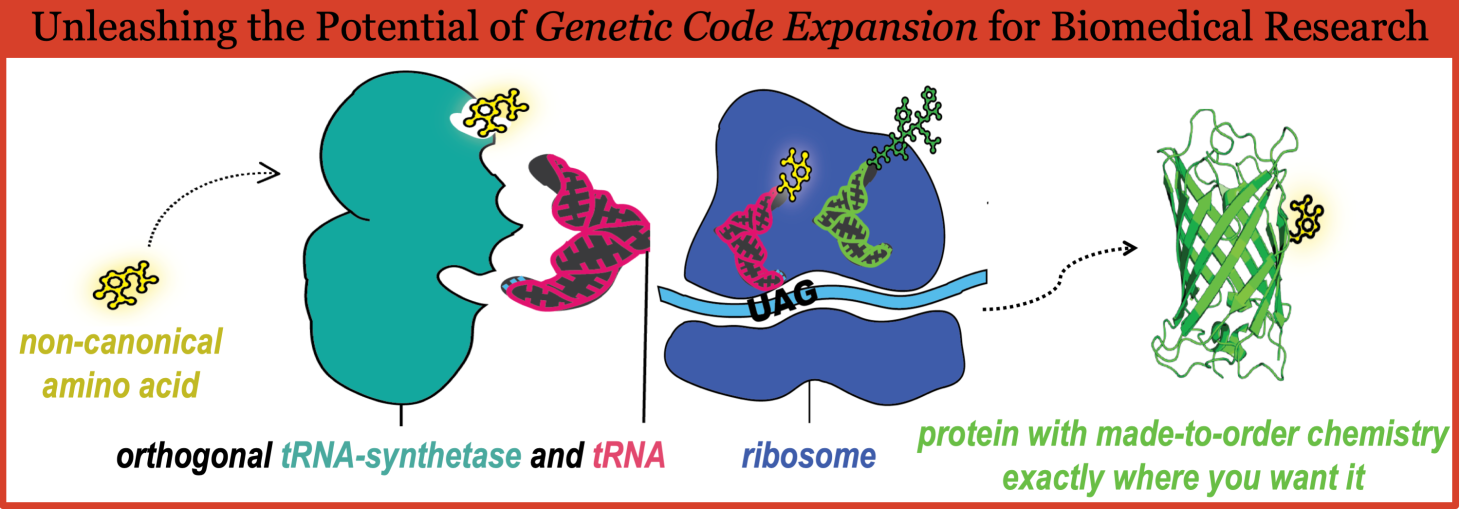
Proteins in all living organisms are made by ribosomes, and the process requires a messenger RNA (mRNA) molecule and a set of transfer RNA (tRNA) molecules that have each been “charged” (i.e. loaded) with its proper amino acid by a corresponding aminoacyl tRNA synthetase (RS) specific for its tRNA and amino acid. The orthogonality of the tRNA charging reactions by all tRNA/RS pairs in cells is essential for maintaining the fidelity of protein translation. Aided by many additional factors, the ribosomes make a protein by having the charged tRNAs – one at a time – recognize their specific “codon” in the mRNA, and in that way, codon-by-codon, the ribosomes translate the nucleic acid sequence of the mRNA to construct a chain of standard amino acid residues in the correct order that was determined by the genetic code.
Genetic Code Expansion (GCE) technology
GCE technology refers to the use of a cell line or organism that contains the addition of a specially engineered “orthogonal” tRNA/RS pair along with a repurposed codon that together allow the encoding of an amino acid that is not among the 20 canonical (i.e. standard) amino acids. This can be done both in prokaryotic cells and in eukaryotic cells.
Since the well-known “universal” genetic code maps 20 common proteinogenic a-amino acids directly to certain codons, the amino acid types that can be incorporated through GCE have been variously referred to as "unnatural", "non-proteinogenic", “non-standard” or "non-canonical". Among these, we prefer the term “non-canonical” amino acid (ncAA) because the amino acids incorporated via GCE approaches can be ones that occur naturally, and through using GCE they have become proteinogenic. The term “orthogonal” above means the engineered tRNA/RS pair does not cross react with any of the 20 canonical amino acids or with the organism’s other RSs or tRNAs.
GCE technology is powerful because it allows for a very wide variety of chemical groups to be incorporated into any target protein at any specified position and has many applications both for basic research and for developing new therapeutics and materials. Some broad areas of GCE applications use:
- ncAAs that have chemical groups with highly selective reactivity that allows them to be covalently attached to other proteins, molecules or surfaces (protein ligations)
- ncAAs that represent naturally-occurring modifications of proteins (i.e. post-translational modifications) enabling more extensive study of their roles in normal regulation or disease,
- ncAAs that have special properties allowing them to be directly used as biochemical probes in a variety of in cell or in vitro studies.
Implementing GCE
If no tools are available for the ncAA or organism of interest, the process can be divided into three main stages:
Constructing the orthogonal genetic components (the tRNA/RS pair) to incorporate the ncAA needed, assembling them in the necessary research organism, and checking that the ncAA-protein is generated. In this stage, it is important that the non-canonical amino acid is not toxic to the cell at the amounts needed, can get into the cell and remain stable and is not recognized by natural tRNA/RS pairs.
Characterizing and optimizing the ncAA expression system. Many variables impact the incorporation efficiency, fidelity and permissivity of the ncAA expression system. Sufficient characterization needs to be completed to trust that your ncAA-protein will be produced as expected under the expression conditions used. Efficiency is the measure of full-length ncAA-protein yield produced compared to wild-type protein. Fidelity is the measure of the ratio of ncAA-protein produced compared to any other type of amino acid residue incorporated. Permissivity is the measure of the variety of other ncAAs that can be incorporated by the new ncAA-tRNA/RS expression system; this quality can be advantageous in that one new tRNA/RS pair can enable the incorporation into proteins of a variety of ncAAs.
Producing the ncAA-protein of interest and studying the structure, interactions, function of the ncAA-protein in vitro or in vivo.
The first two steps currently present a big barrier hindering access to GCE by research labs that would greatly benefit from using it. To address this problem, a major thrust of the GCE4All Center is to develop robust tools and protocols for ncAAs and organism of wide interest. Furthermore, an additional thrust – to be pursued in parallel – is to develop robust processes for the Stage 1 and Stage 2 activities; these will make it possible for any molecular biology group to do what is needed to implement GCE for a new ncAA.
Learn more about GCE
The above is only a very brief overview of the basics of GCE. For those seeking more extensive information about GCE and its applications, a good place to start is a 2021 review article entitled Genetic Code Expansion: Inception, Development, Commercialization. Another up-to-date resource is a complete issue of the Journal of Molecular Biology that focuses on recent advances in Genetic Code Expansion: from cell engineering to protein design.